The Jmol Window and Console

Jmol is a free web-based program that displays protein structures in a fully interactive 3-dimensional browser window.
On the right side of the screen is a live Jmol window showing an example protein called a zinc finger motif.
Click and drag with the left mouse button to rotate this display. Click and drag with the left mouse button while holding shift to zoom in and out.

On the bottom right, below the Jmol window, is the Jmol Console. This is where text commands can be entered to alter the way a protein is displayed in Jmol, such as changing the protein's colors and display formats.
Changing Colors with the Jmol Console
When a protein structure is first displayed in Jmol, it will show every atom colored by element (carbon is gray, oxygen is red, nitrogen is blue).
While visually impressive, this default display can also be visually overwhelming. It is hard to see the path of the protein chain(s), identify secondary structures or locate individual amino acid sidechains.Changing the color scheme can begin to clarify these features.
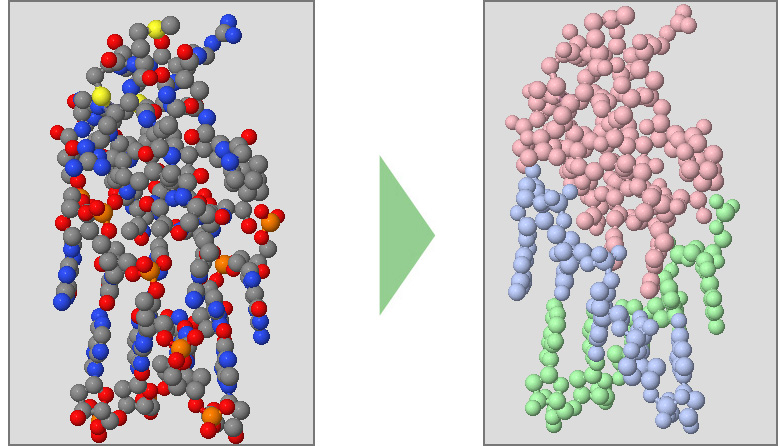

Below are a collection of Jmol color commands.
When entered into the Jmol console, these commands will alter the Jmol display. You are strongly encouraged to practice entering these commands into the console yourself to become familiar with how they work.
Clicking the Watch this Command Demonstrated buttons will also demonstrate each command on the protein shown in the Jmol window.
-
Color Chains will make each protein strand (often referred to as "chains") a different color.
color chains
Watch This Command Demonstrated -
Color Structure will highlight the secondary structures in your protein. Alpha Helices are colored magenta, Beta Pleated Sheets are colored yellow, and Nucleic Acids (DNA and RNA) are colored purple.
color structure
Watch This Command Demonstrated -
Color CPK will color each atom by what element it is (carbon is gray, oxygen is red, nitrogen is blue). This is the default color scheme applied to new structures when first opened.
color cpk
Watch This Command Demonstrated -
Specific Colors can also be applied to your protein design. This command can use common color names or RGB color values.
color orange
Watch This Command Demonstratedcolor [150, 25, 225]
Watch This Command Demonstrated
The color commands listed above are only a small portion of the Jmol color commands that can be entered into the console. For a comprehensive list of Jmol color commands, visit http://jmol.sourceforge.net/jscolors/.
Key color commands are summarized on the Jmol Quick Reference Sheet, which is also linked to in the left column of this webpage.
Changing Display Formats with the Jmol Console
Jmol color commands can help visually simplify a protein structure or draw attention to specific features of the molecule, such as the secondary structures.
In a similar way, the Jmol display format commands can be used to clarify features of a protein structure.
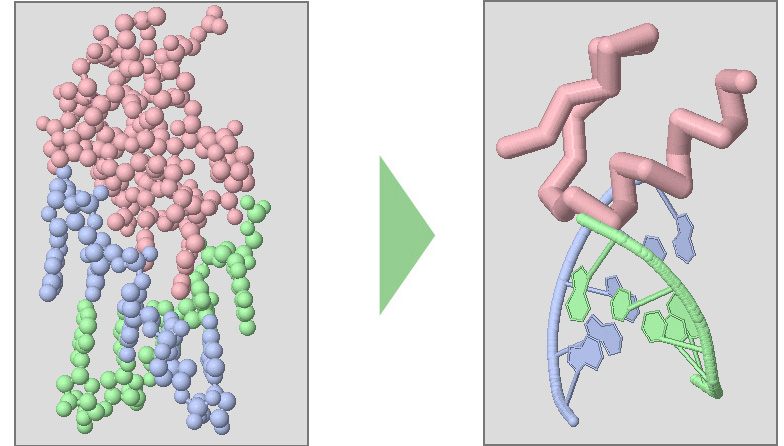

Below are a collection of Jmol display format commands.
When entered into the Jmol console, these commands will alters the Jmol display. Clicking the Watch this Command Demonstrated buttons will also demonstrate each command on the protein shown in the Jmol window.
-
Wireframe Format displays all of the covalent bonds in a protein structure as thin lines, and is a great way to see how atoms are connected. It can, however, be visually overwhelming and difficult to see the path of a protein chain.
wireframe only
Watch This Command DemonstratedYou can also use a number to size your wireframe lines to an exact diameter (in angstroms). Make sure to use a decimal in your number or your command will not work properly (ie: use "wireframe 1.0", not "wireframe 1").
wirefame 0.5
Watch This Command DemonstratedYou can also completely remove a display format by adding off to your wireframe command. Note that this does not permanantly delete atoms from your molecular structure file. Rather, you are just turning off any sort of visual representation of them, meaning they can be added back later.
wirefame off
Watch This Command Demonstrated -
Spacefill Format displays atoms as large spheres, and is a great way to see a protein's overall globular shape. Like Wireframe Format, it can be difficult to see the path of a protein backbone with spacefill format.
spacefill only
Watch This Command Demonstratedspacefill 1.0
Watch This Command Demonstratedspacefill off
Watch This Command Demonstrated -
Backbone Format displays only the alpha carbon atom of each amino acid in a protein structure, and connects them with a line. It is a great way to see the overall fold of amino acid strands in a protein structure, but does not display the sidechain atoms.
backbone only
Watch This Command Demonstratedbackbone 1.25
Watch This Command Demonstratedbackbone off
Watch This Command Demonstrated -
Cartoon Format is similar to backbone format, in that it represents only the backbon e path of a protein chain. It is a great way to see the overall fold of amino acid strands in a protein structure, but does not display the sidechain atoms. Cartoon format also shows DNA base pairs.
cartoon only
Watch This Command Demonstratedcartoon off
Watch This Command Demonstrated -
Multiple Display Formats can be applied to a protein structure at the same time. This allows you to combine wireframe, spacefill and backbone elements into one design.
Each of the three commands demonstrated below would be submitted to the console separately, by either clicking "execute" or the "enter" key.
spacefill 0.45
Watch This Command Demonstrated
wireframe 0.2
backbone 1.25
The display format commands listed above are only a small portion of the Jmol dispaly format commands that can be entered into the console. For a comprehensive list of Jmol display format commands, visit https://chemapps.stolaf.edu/jmol/docs/.
Key display format commands are summarized on the Jmol Quick Reference Sheet, which is also linked to in the left column of this webpage.
Part 1 - Conclusion
Jmol is an open-source molecular visualization program that allows 3-dimensional protein structures to be displayed in an interactive browser window.
Jmol color and display format commands can be entered into the Jmol Console to change the appearance of the protein structure being visualized. Different color schemes and display formats highlight different features of the protein
Take time to practice the commands introduced in this section of the Protein Modeling Event Jmol Training Guide. When ready, move on to part 2, which will introduce the select command.
