The Select Command
The color and display formats commands can provide a feel for the size, number of chains, and overall shape of an entire protein structure.
However, because most proteins are quite large, each year's prebuild model will not represent an entire protein structure, but rather a subset of the amino acids in a larger protein.
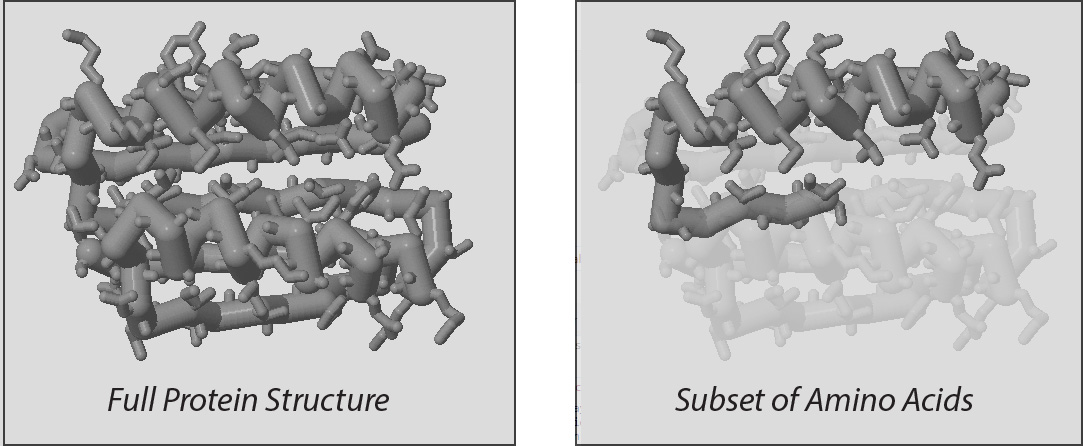

Up until this point, every Jmol command has impacted every amino acid in the molecular structure being displayed. But with the select command, you can control what subset of amino acids future Jmol commands will affect.
An example protein called Top 7 will be used to demonstrate the select commands that are introduced below. It will be colored gray and displayed with backbone 1.25 and wireframe 0.5.
For each select command, the command color magenta is also applied in order to draw attention to what atoms were selected.
-
Select All will make all future commands impact every atom in the structure. This is the default selection when a structure is first opened in Jmol.
select all
Watch This Command Demonstrated
color magenta
Note: If you want to return the protein structure being displayed to the right back to the original gray color scheme, enter the command select all followed by color gray.
-
Select Backbone will make all future commands only impact atoms that are part of your protein backbone.
select backbone
Watch This Command Demonstrated
color magenta -
Select Sidechain will make all future commands only impact atoms that are part of your protein's sidechains.
select sidechain
Watch This Command Demonstrated
color magenta -
Select Helix will make all future commands only impact atoms that are part of alpha helix secondary structures.
select helix
Watch This Command Demonstrated
color magenta -
Select Sheet will make all future commands only impact atoms that are part of beta pleated sheet secondary structures.
select sheet
Watch This Command Demonstrated
color magenta -
Select by Residue Number will make all future commands only impact atoms that are part of a specific amino acid or nucleic acid monomer. Each amino acid or nucleic acid in a structure file is given a residue number, which can be used to select that residue in Jmol.
select 82
Watch This Command Demonstrated
color magenta -
Select by Residue Range will make all future commands only impact atoms that are part of a specific amino acid or nucleic acid residue range.
select 60-82
Watch This Command Demonstrated
color magenta -
Select Atom Number will make all future commands impact only a single atom in the structure. Each atom in a structure file is given a unique atom number, which can be used to select that atom in Jmol. To differentiate atom numbers from residue numbers, a "@" symbol is added at the beginning of the atom number.
select @595
Watch This Command Demonstrated
color magenta -
Select by Element will make all future commands impact only atoms of a specific element type (ie: carbon, nitrogen, oxygen, etc.).
select nitrogen
Watch This Command Demonstrated
color magenta -
Select by Amino Acid Type will make all future commands impact only atoms that are part of a specific amino acid type (ie: histidine, cysteine, arginine, etc.). Note that this command does not use the full amino acid names, but rather their three-letter abbreviations.
Click Here for a full list of the amino acid abbreviations on page 2 of the Jmol Quick Reference Sheet.
select thr
Watch This Command Demonstrated
color magenta -
Select by Chain ID will make all future commands only impact atoms that are part of a specific chain. Each protein or nucleic acid chain in a structure file is given a unique identifier (usually a single letter), which can be used after a ":" to select that chain in Jmol.
Note: The "Top 7" molecular structure only has one protein chain. So for this example, the four chain protein "Hemoglobin" will be used. Click Here to load the hemoglobin structure.
select :C
Watch This Command Demonstrated
color magenta

Zero Atoms Selected!
Every time you use the select command, the number of atoms you have selected will be displayed in the console.
But if you have submitted a select command that does not include any atoms, it will display "0 atoms selected" in the console. For example, our "Top 7" protein is only 94 amino acids long, so "select 100" would return the message "0 atoms selected".

Identifying Atoms
Often you may not know the exact amino acid type, residue number, element or atom number of something you want to select.
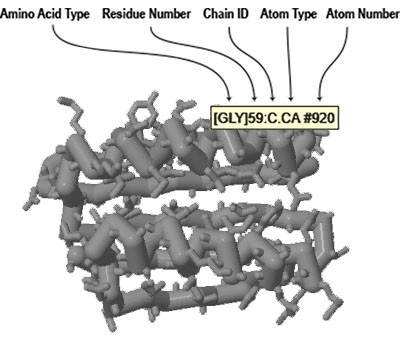
The easiest way to determine this information is to roll your mouse over the atom you want to identify and wait for a moment until a small yellow information box appears. This box will include all of the information needed to select the atom being hovered over.

The select commands listed above are only a small portion of the Jmol select commands that can be entered into the console. For a comprehensive list of Jmol display format commands, visit https://chemapps.stolaf.edu/jmol/docs/.
Key select commands are summarized on the Jmol Quick Reference Sheet, which is also linked to in the left column of this webpage.
Boolean Operators
Even with the predefined selection types discussed above, it can still be difficult to select an exact subset of atoms. The boolean operator commands allow for even more nuanced selection types.
To understand how boolean operators work, imagine working with two predefined selection types:helix and backbone. Some atoms in our "Top 7" protein are part of helical secondary structures, some atoms are part of the protein backbone, are some atoms would be in both selection types.
-
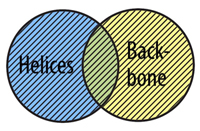
The OR Command allows you to select atoms from more than one selection type. In order for an atom to be selected using or, it does not need to fit into both selection types at the same time, just one or the other.
select helix or backbone
Watch This Command Demonstrated
color magenta -
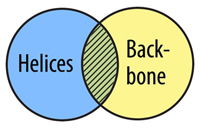
The AND Command allows you to select atoms that are in two selection types. In order for an atom to be selected using and, it must fit into both selection types at the same time, not just one or the other.
select helix and backbone
Watch This Command Demonstrated
color magenta
Part 2 - Conclusion
The Select Command allows future Jmol commands to only impact a subset of atoms in a larger protein structure.
Boolean Operators allow for even more refined selection commands by grouping more than one selection type together with "and" or "or" commands.
Take time to practice the commands introduced in this section of the Protein Modeling Event Jmol Training Guide. When ready, move on to part 3, which will introduce adding sidechains to a backbone model.
